Exploring evolutionary adaptations in organisms lacking a conventional respiratory chain
Many bacteria and lower eukaryotes have reconfigured or are completely devoid of a conventional electron transport chain (ETC) such as that found in mammalian mitochondria. We are interested in systematically exploring the evolutionary adaptations that allow organisms to tolerate missing components of the ETC. Such evolutionary innovations are attractive for several reasons: 1) they can inspire novel strategies of “bypassing” a corrupted ETC in various models of mitochondrial disorders; 2) they can provide a basis for developing tools to dissect cellular redox metabolism; and 3) they can offer insight into how electron chains composed of multiple redox cofactors control electron flow and enable catalysis.
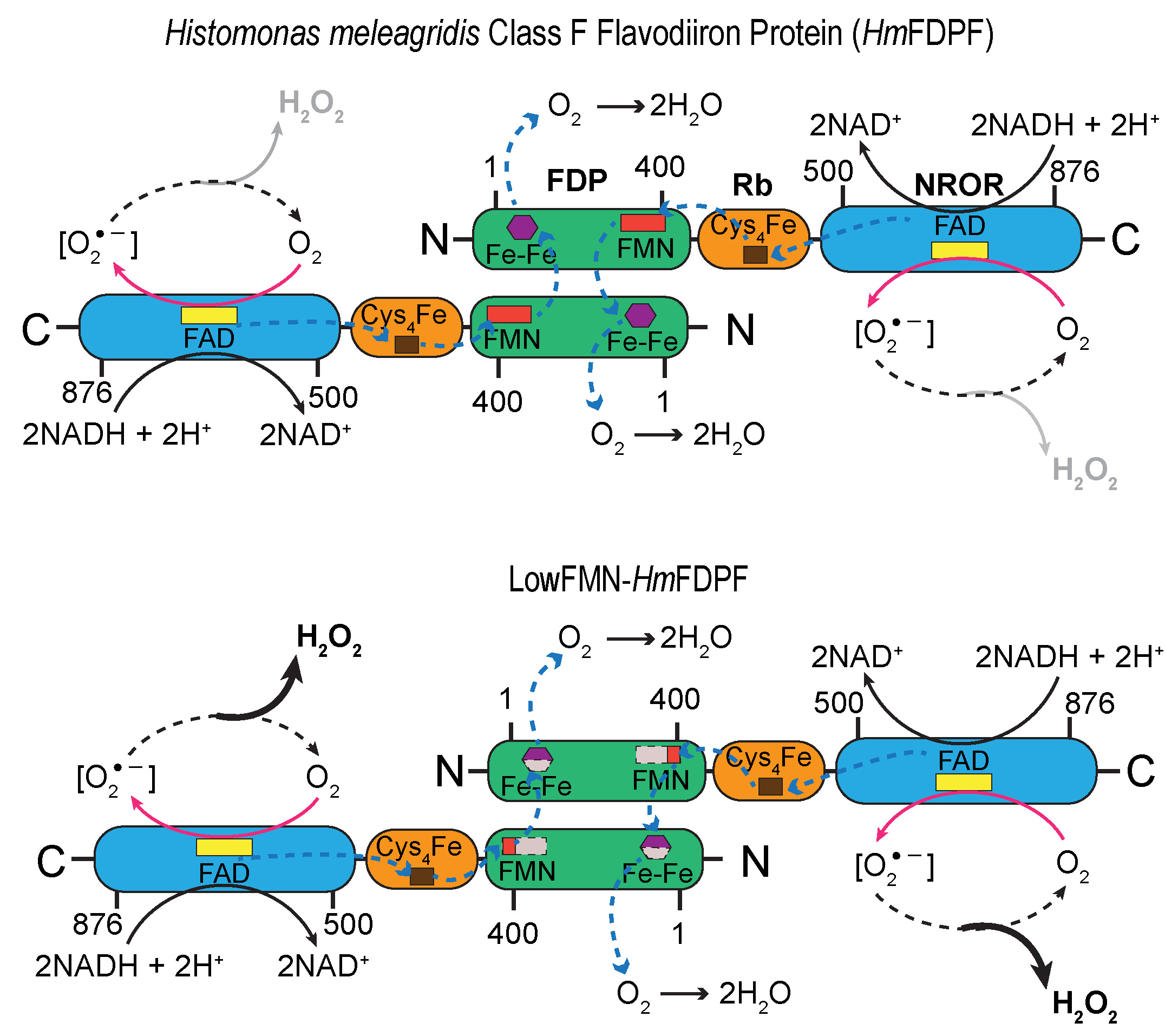
For example, a number of bacteria and members of the Excavata supergroup of unicellular eukaryotes can tolerate fairly aerobic environments despite lacking a conventional ETC, which is ordinarily thought to play a pivotal role in detoxifying environmental oxygen. These organisms rely on enzymes such as H2O-forming NADH oxidases (NOXes) and flavodiiron proteins (FDPs), which catalyze a H2O-forming NADH oxidase reaction to recycle NAD+ using oxygen as the final electron acceptor. Our recent studies provided important insight into how multiple redox components within a multidomain terminal oxidase FDP support electron transport without substantial side reactions.
Abdulaziz EN, Bell TA, Rashid B, Heacock ML, Begic T, Skinner OS, Yaseen MA, Chao LH, Mootha VK, Pierik AJ, Cracan V. (2022) A natural fusion of flavodiiron, rubredoxin, and rubredoxin oxidoreductase domains is a self-sufficient water-forming oxidase of Trichomonas vaginalis. J. Biol. Chem. Aug;298(8):102210 hhtps://www.jbc.org/article/S0021-9258(22)00652-4/fulltext
Munan S, Yoval-Sánchez B, Yao C, Seravalli J, Abdulaziz EN, Heacock ML, Ramesh A, Galkin A, Cracan V. (2025). Biochemical characterization of a flavodiiron protein from bird parasite Histomonas meleagridis: superoxide as a reaction intermediate. J. Biol. Chem. Sep 4:110685. https://www.jbc.org/article/S0021-9258(25)02537-2/fulltext
Development of genetically-encoded tools for redox signaling research

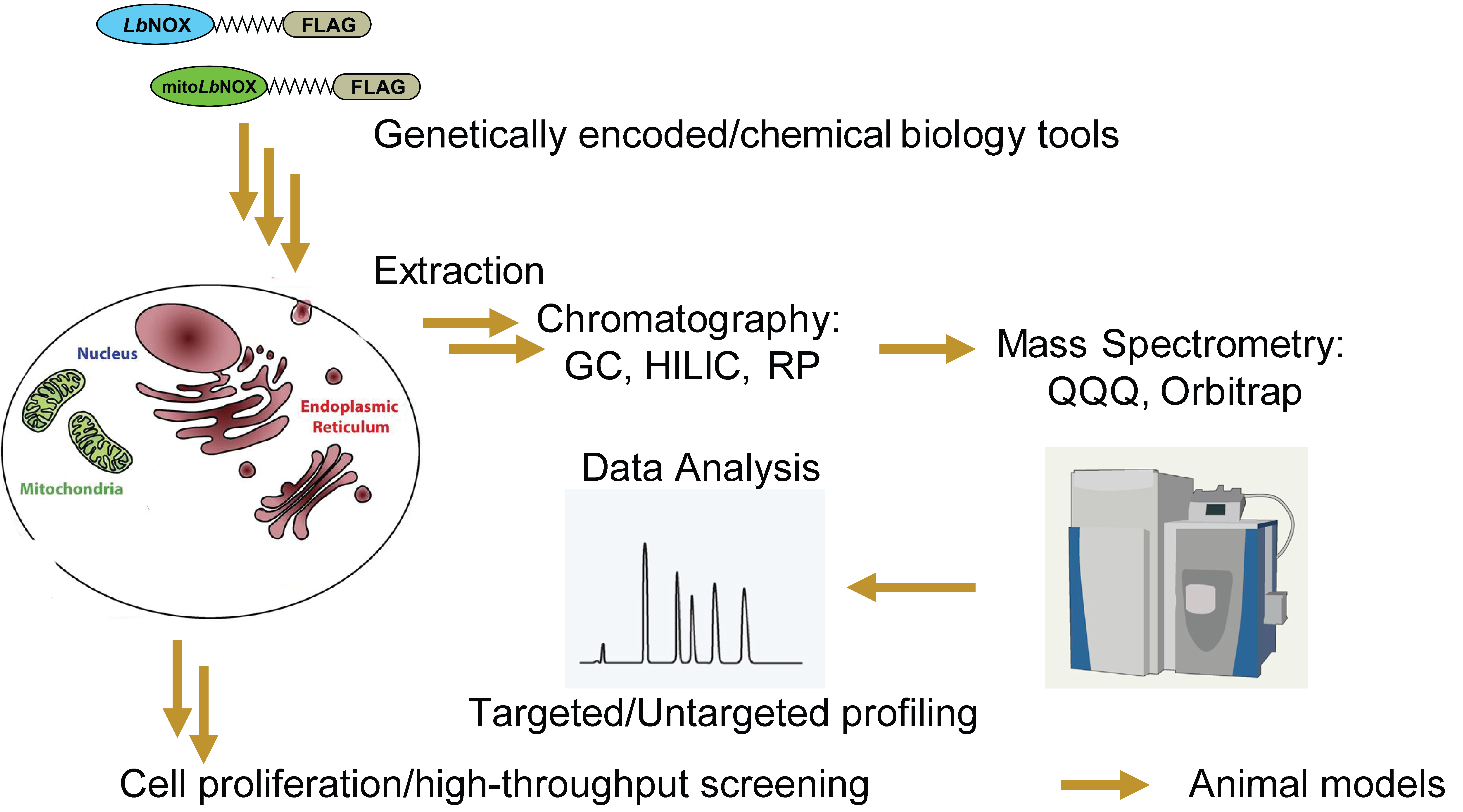
The coenzymes nicotinamide adenine dinucleotide (NAD) and nicotinamide adenine dinucleotide phosphate (NADP) serve as electron donors and acceptors in a wide variety of metabolic reactions and are thus major contributors to the cellular redox environment. Engineering tools to perturb or visualize the redox state of these coenzyme pools would enable studies of the role of redox imbalances in driving numerous pathologies. Previously, we used a H2O-forming NADH oxidase from the bacterium Lactobacillus brevis (LbNOX) to ameliorate the proliferative defect caused by impaired ETC in human cells (Titov, Cracan et al. Science; 2016). If the organ pathologies associated with mitochondrial diseases or other conditions stem from redox imbalance, then expression of LbNOX or similar proteins holds promise as a “protein prosthesis” therapy. Based on LbNOX, we subsequently engineered a novel H2O-forming oxidase which is strictly specific towards NADPH (TPNOX), enabling direct, compartment-specific decrease in NADPH/NADP+ ratio in mammalian cells (Cracan et al. Nature Chem Bio; 2017). We demonstrated using LbNOX and TPNOX that reducing equivalents do not rapidly exchange between NADH/NAD+ and NADPH/NADP+ in the mammalian cytosol, but that they do exchange readily in the mitochondria.
More recently, we introduced a new genetically encoded tool that is based on a soluble transhydrogenase from E. coli (EcSTH) to directly increase cellular NADH/NAD+ ratio with a compartment specific precision. Another tool we developed is based on a di-domain glycerol-3-phosphate dehydrogenase from algae Chlamydomonas reinhardtii (CrGPDH), which alleviates metabolic consequences of the NADH-reductive stress under hypoxic conditions. We aim to further expand our toolkit of genetically encoded tools to ultimately tease apart the logic of redox compartmentalization in different backgrounds and conditions.
Parkhitko AA and Cracan V. (2025) Xenotopic synthetic biology: Prospective tools for delaying aging and age-related diseases. Science Advances. https://www.science.org/doi/10.1126/sciadv.adu1710
Pan X, Heacock ML, Abdulaziz EN, Violante S, Zuckerman AL, Shrestha N, Yao C, Goodman RP, Cross JR, Cracan V. (2024) A genetically encoded tool to increase cellular NADH/NAD+ ratio in living cells. Nat. Chem. Biol. https://www.nature.com/articles/s41589-023-01460-w
Pan X, Munan S, Zuckerman AL, Pon A, Violante S, Cross JR, Shah H and Cracan V. (2025). A genetically encoded bifunctional enzyme mitigates redox imbalance and lipotoxicity via engineered Gro3P-Glycerol shunt. bioRxiv. 2025.06.02.657195 https: //doi.org/10.1101/2025.06.02.657195
Understanding how metabolism is rewired in cancer
It has become increasingly clear that, in order to meet their unique proliferative needs, cancer cells exhibit altered metabolism compared to normal cells-so called “metabolic rewiring.” For example, escalated reactive oxygen species (ROS) generation by cancer cells has been recognized as an important contributor of tumor development and progression. Our knowledge of how cancer cells maintain the delicate balance between ROS production and detoxification is still very limited. We and others established that redox requirements are important for proliferation in cancer cells (Titov, Cracan et al. Science; 2016). Using a combination of genetically encoded tools and metabolomics we aim to better understand how different cancers rewire their redox metabolism based on tissue of origin or cell type. We are ultimately interested in harnessing the differences in redox metabolism between cancer and normal cells to develop novel therapeutic interventions.